RNA at the Center of Epigenetics
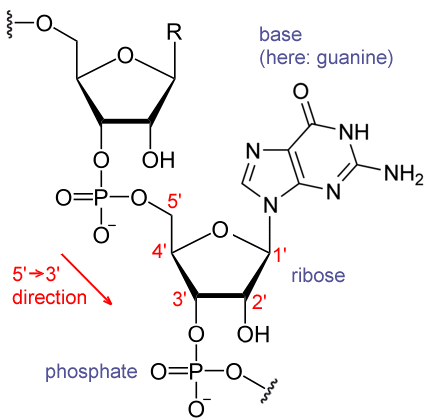
Researchers from Tel Aviv University, Sheba Medical Center, and University of Chicago have discovered a new RNA modification that plays a central role in the regulation of gene expression. The methylation of Adenosine in position 1 occurs on thousands of different transcripts in eukaryotic cells and in 20% of human transcripts. The presence of the methyl group near the translation start site correlates with protein production. The findings were published in the journal Nature.
According to the classical definition, epigenetics regulates gene expression through chemical modifications –mainly methylations and adenylations- in DNA and proteins like histones. Depending on the organism, the modified protein residue or the type of modification, the resulting DNA structure favors or hinders gene expression. Four years ago, Prof. Rechavi discovered that some mRNA adenines were methylated in position six (m6A). The modification affected specific regions of the transcript, was detected by specific proteins and responded to environmental cues. The discovery of the RNA methylation affecting the trancripts stability, localization and translability inaugurated the field of epitranscriptomics.
New RNA adenosine modification
In their last study, the team led by Prof. Rechavi discovered the methylation of mRNA adenosine in position 1 (m1A). The modification was found in structured regions near canonical and non-canonical translation initiation sites, and is linked to protein synthesis.
The discovery of the new epigenetic mark on RNAs opens the door to a deeper understanding of gene expression and cellular processes. The researchers are studying the possible relationship between m1A disruption and cancer and neurodegenerative disease.
Source: AFTAU